Image Classification and Convolutional Neural Networks
Over the past few years, deep learning techniques have dominated computer vision. One of the computer vision application areas where deep learning excels is image classification with Convolutional Neural Networks (CNNs).
The goal of image classification is to classify a specific image according to a set of possible categories. State-of-the-art image classifiers often result from transfer learning approaches based on pre-trained convolutional neural networks.
In this article, we illustrate the training of a plant disease classification model using the Fastai library.
The Fastai Library
The fastai library simplifies and enables the training of fast and accurate neural nets using modern best practices. It’s an excellent initiative by Jeremy Howard and his team aimed at democratizing deep learning and making it easier for everyone to build deep learning models.
Fastai is a wrapper for PyTorch, which makes it easier to access recommended best practices for training deep learning models, while at the same time making all the underlying PyTorch functionality directly available to developers.
The main reason why fastai has been successful is the use of transfer learning techniques. Transfer learning involves the use of a pre-trained model, that has been trained on a large collection of data and finally repurposed for ones’ specific needs. Fastai library has awesome documentation to help you navigate through their library functions with live examples on how to use them.
Training a Plant Disease Classifier
The Dataset
The data used in this article is obtained from the PlantVillage Disease Classification Challenge organized by CrowdAi. The goal of this challenge was to develop algorithms that can accurately diagnose a disease based on a plant image. PlantVillage is a not-for-profit project by Penn State University in the US and EPFL (Ecole polytechnique fédérale de Lausanne) in Switzerland.
The organizations empower smallholder farmers to increase yield by leveraging Artificial Intelligence to provide offline expert level knowledge and extension advice. PlantVillage has already collected and continue to collect tens of thousands of images of diseased and healthy crops.
The same dataset of diseased plant leaf images and corresponding labels comprising 38 classes of crop disease can also be found in spMohanty’s GitHub account.
Training the Model
We use the vision module of the Fastai library to train an image classification model which can recognize plant diseases at state-of-the-art accuracy. While training of the model can be done locally using a laptop, we use Google Colab which gives us more compute power, access to a GPU, and an easy-to-use Jupyter notebook environment for building machine learning and deep learning models.
We begin by placing the following three lines at the start of the notebook to ensure that any edits made to libraries are reloaded automatically, and also that any charts or images displayed are displayed within the notebook. These are not Python codes but special directives for Jupyter Notebook itself. The % is one of the magic commands supported in Jupyter Lab that adds extra functionality to our Jupyter Lab notebooks (and isn’t limited to the core language).
The next step is to import the required libraries. The fastai module and any other module can easily be installed using the pip command.
Loading and looking at the Data
We download the colored (original RGB) images using the following command:
! svn export https://github.com/spMohanty/PlantVillage-Dataset/trunk/raw/colorWhenever we approach a problem, the first thing to do is to take a look at the data in order to better understand what the problem is and what the data looks like before we can figure out how to solve the problem. Taking a look at the data means understanding how the data directories are structured, what the labels are, and what some of the sample images look like.
In this particular dataset, the folder name represents the class label of all the images present within that folder. We need to extract the label names from the folder name automatically. Fortunately, fastai library provides the ImageDataBunch.from_folder function that enables automatic extraction of the label names from the folder name. In addition, the ImageDataBunch class makes it easy to create the training and validation sets with images and labels. Once the data is loaded, we can also normalize the data by using .normalize to ImageNet parameters.
np.random.seed(8)
bs = 64
tfms = get_transforms(flip_vert=True, max_warp=0)
data = ImageDataBunch.from_folder(path_train,
valid_pct=0.2,
train=".",
# test="../test images",
ds_tfms=tfms,
size=224,bs=bs,
num_workers=0).normalize(imagenet_stats)The .show_batch() function of the ImageDataBunch class can be used to view a random sample of images from the given data.
data.show_batch(rows=3, figsize=(10,8))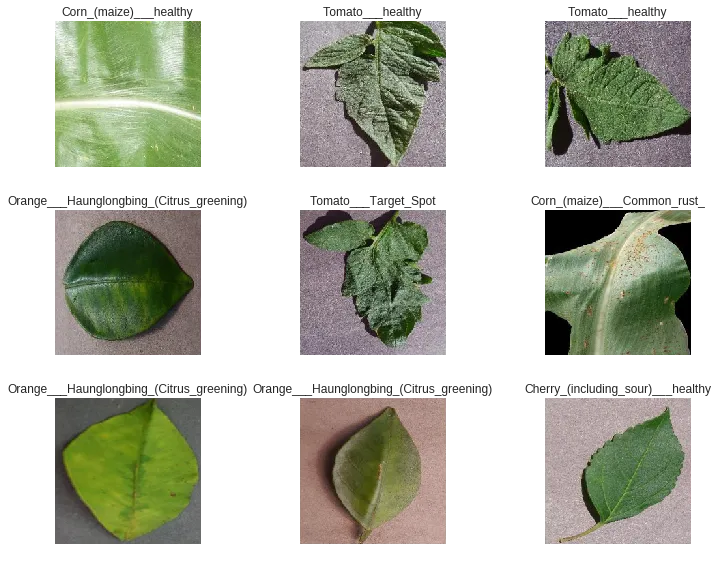
You’ll notice that the images appear to have been zoomed and cropped in a reasonably nice way. Fastai provides a rich image transformation library, whose main purpose is data augmentation when training computer vision models. The library can, however, be used for other general transformation tasks such as default center cropping.
DataBunch has a property called c which helps determine the number of classes—an important piece of information, especially for classification problems. We print the class labels and the number of classes as follows:
print(data.classes)
len(data.classes),data.c
Transfer learning using a Pre-trained model: ResNet 50
Now that our data is ready, it’s time to fit a model. To create the model we will use the function create_cnn from Learner class and feed a pre-trained model, in this case, ResNet 50, from the models class. The model we are building will take images as input and outputs the predicted probability of each category, with accuracy being used as the measure of performance.
arch = models.resnet50
learn = create_cnn(data=data,arch=arch, metrics=accuracy)We use the method fit_one_cycle to train the model for 4 epochs (4 cycles through the data).
learn.fit_one_cycle(4)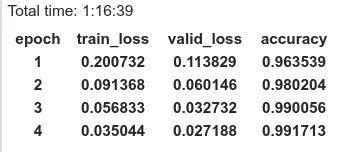
The set of weights generated from the training process can be saved for re-use using the .save function of the learner class. In our case, we save it as ‘stage-1’.
learn.save('stage-1')Conclusion
To achieve global food security, society needs to increase food production by an estimated 70% in order to feed an expected population size of over 9 billion people by 2050.
One way of achieving this target is to reduce infectious crop diseases, which often result in a reduction of the potential yield by an average of 40% and sometimes even more—especially for small-scale farmers in the developing world.
The widespread adoption of smartphones around the globe provides the potential of turning our phones into valuable disease diagnostics tools, enabling farmers to recognize diseases from images captured using the device’s camera.
Discuss this post on Hacker News and Reddit.

Comments 0 Responses