Many packages in Python also have an interface in R. Keras by RStudio is the R implementation of the Keras Python package. Most of the functions are the same as in Python. The only difference is mostly in language syntax such as variable declaration.
In this tutorial, we’ll use the Keras R package to see how we can solve a classification problem. We’ll use the Kyphosis dataset to build a classification model. Kyphosis is a medical condition that causes a forward curving of the back—so we’ll be classifying whether kyphosis is present or absent.
We’ll get started by started by loading in the TidyVerse library that will provide us with the function to read in the dataset.
library(tidyverse)We’ll now import Keras so that we can access the Keras functions then read in the Kyphosis dataset.
library(keras)
df <- read.csv('kyphosis.csv')We can quickly check the head in order to get a glimpse into the dataset that we’re working with.
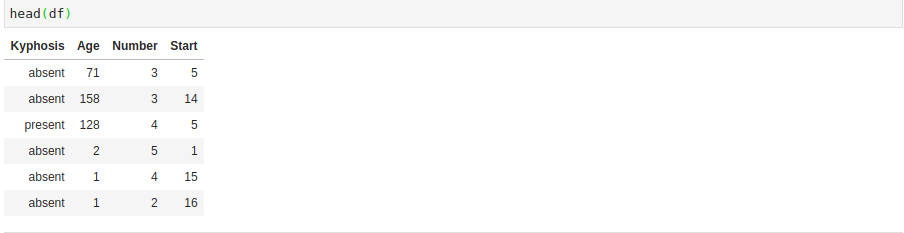
We’ll be working to predict the Kyphosis column. The first thing we’ll do is convert the Kyphosis column into zeros and ones since the machine learning model will only accept numerical figures. In order to achieve this, we’ll use the fastDummies library that will help us in creating the dummy variables. In the process, we’ll remove the first dummy variable to avoid the dummy variable trap, as we have seen in previous machine learning tutorials. After that, we’ll check to make sure the head of the data that has been generated.
library(fastDummies)
dummy_data <- fastDummies::dummy_cols(df,remove_first_dummy = TRUE)
head(dummy_data) 
We can see that the previous Kyphosis column is still in our dataset. We’ll now work to remove it so that we’re left only with the one we just generated. After that, we can then check its head.
keep <- c('Age','Number','Start','Kyphosis_present')
final <- dummy_data[keep]
head(final)
We’re going to use the Caret library to prepare the data for our classification model. We’ll first create the index that we shall use to split the data into a training and testing set.
library(caret)
index <- createDataPartition(final$Kyphosis_present, p=0.7, list=FALSE)
We’ve created an index that will use 70% of the data on training and the other 30% for a test set.
final.training <- final[index,]
final.test <- final[-index,]The next step is to prepare the data for Keras by scaling it. We’ll use the scale function to achieve this. We’ll then use the to_categorical function on the labels to ensure that they’re categorical.
X_train <- final.training %>%
select(-Kyphosis_present) %>%
scale()
y_train <- to_categorical(final.training$Kyphosis_present)We’ll perform a similar transformation on the test set and the test set labels.
X_test <- final.test %>%
select(-Kyphosis_present) %>%
scale()
y_test <- to_categorical(final.test$Kyphosis_present)The next step is to define our model. We’ll use keras_model_sequential() to initialize the model. This is similar to the Sequential function in Keras Python. We’ll then define dense layers using the popular relu activation function.
We also add drop-out layers to fight overfitting in our model. Similar to Keras in Python, we then add the output layer with the sigmoid activation function.
The next step is to compile the model using the binary_crossentropy loss function. This is because we’re solving a binary classification problem.
We’ll use the adam optimizer for gradient descent and use accuracy for the metrics. We then fit our model to the training and testing set. Our model will run on 100 epochs using a batch size of 5 and a 30% validation split.
model <- keras_model_sequential()
model %>%
layer_dense(units = 256, activation = 'relu', input_shape = ncol(X_train)) %>%
layer_dropout(rate = 0.4) %>%
layer_dense(units = 128, activation = 'relu') %>%
layer_dropout(rate = 0.3) %>%
layer_dense(units = 2, activation = 'sigmoid')
history <- model %>% compile(
loss = 'binary_crossentropy',
optimizer = 'adam',
metrics = c('accuracy')
)
model %>% fit(
X_train, y_train,
epochs = 100,
batch_size = 5,
validation_split = 0.3
)We can now look at our model’s summary.
summary(model)The next thing we can do is evaluate the model. We’ll check the training loss and its accuracy.
model %>% evaluate(X_test, y_test)We can also visualize these results using a graph. We’ll start with visualizing the model loss.
plot(history$metrics$loss, main="Model Loss", xlab = "epoch", ylab="loss", col="orange", type="l")
lines(history$metrics$val_loss, col="skyblue")
legend("topright", c("Training","Testing"), col=c("orange", "skyblue"), lty=c(1,1))
The model accuracy can be visualized in a similar manner.
plot(history$metrics$acc, main="Model Accuracy", xlab = "epoch", ylab="accuracy", col="orange", type="l")
lines(history$metrics$val_acc, col="skyblue")
legend("topleft", c("Training","Testing"), col=c("orange", "skyblue"), lty=c(1,1))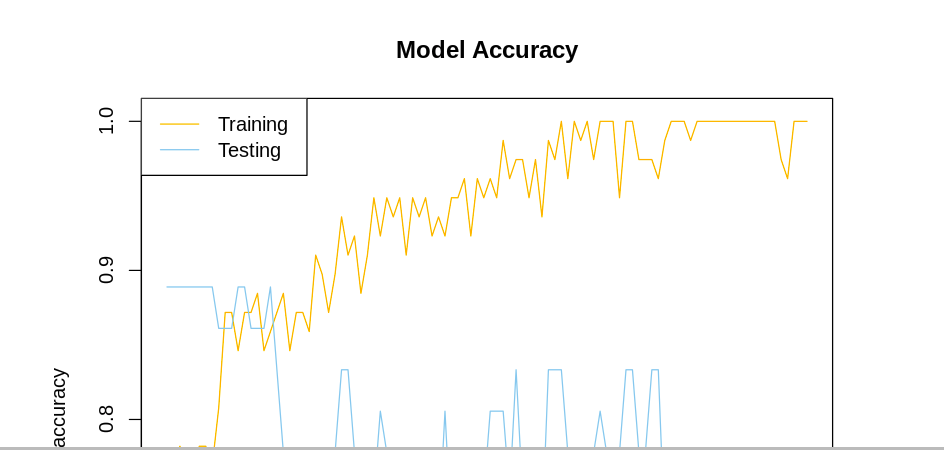
From the model loss graph, we notice that the training loss goes down but the testing loss is up. We’d have to tune the parameters further to ensure the testing loss is decreasing, too. In the model accuracy we notice that the training accuracy is also higher than the testing accuracy. Let’s now move forward to make predictions using the predict_classes Keras function.
predictions <- model %>% predict_classes(X_test)
At this point, we can print the confusion matrix. This will help us see how many predictions are being made correctly, and otherwise.
table(factor(predictions, levels=min(final.test$Kyphosis_present):max(final.test$Kyphosis_present)),factor(final.test$Kyphosis_present, levels=min(final.test$Kyphosis_present):max(final.test$Kyphosis_present)))
At this point, we might be interested in saving our model for future use. In order for this to happen, we need to install h5py. R Keras will use it to save the model.
conda install h5pyLet’s now proceed to show how we can save the model and load it.
save_model_hdf5(model, "rkerasmodel.h5")
model <- load_model_hdf5("rkerasmodel.h5")R Keras allows us to build deep learning models just like we would using Keras in Python. This makes it very easy for someone who has used Keras in any language to transition smoothly between other languages. You can learn more about R Keras from its official site.

Comments 0 Responses