Quick Overview
This tutorial documents a GitHub project called GARI (Genetic Algorithm for Reproducing Images). The project is available here:
Before discussing the details of the project, let’s run through a quick overview of it.
The GARI project accepts an image as input. This image can have one or more channels (i.e. the image could be binary, gray, or color, such as RGB). RGB is the most popular color model that produces any color as a combination of the 3 color channels Red, Green, and Blue. Hence its abbreviation.
The genetic algorithm (GA) starts from a randomly generated image of the same shape as the input image. This randomly generated image is evolved, using crossover and mutation, using GA until it reproduces an image similar to the original one. The exact original image might not be accurately reproduced, but at least a similar image will be generated.
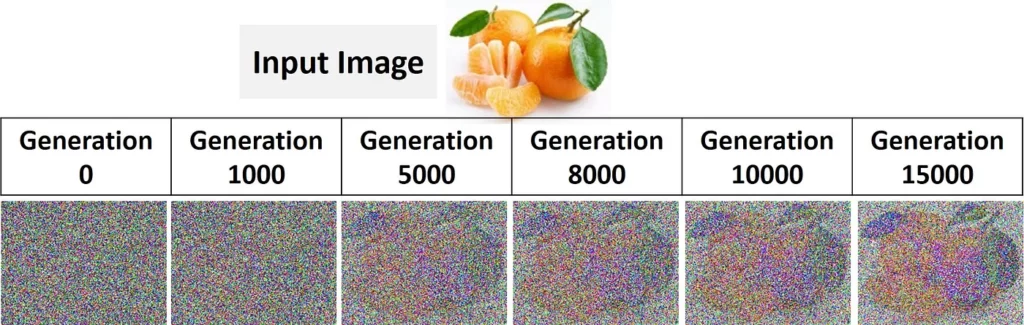
The project consists of 2 Python files. The first one is named “GARI.py” which holds the implementation of the GA functions responsible for reproducing the image. The other file is named “Example_GeneticAlgorithm.py”, which just calls the functions defined in the previous file. It reads a colored image file named “fruit.jpg”, available in the project, and passes it through the functions of GA. The next figure shows that image and how it’s reproduced by GARI after 15,000 generations.

Further Reading about GA and its Python Implementation
Before going further in this tutorial, I’d recommend that you establish a solid background on how GA works. I prepared a number of resources that might help you to start:
- [Book] “Ahmed Fawzy Gad ‘Practical Computer Vision Applications Using Deep Learning with CNNs’. Dec. 2018, Apress, 978–1–4842–4167–7 “. I covered GA in one of the chapters of this book. GA is used for optimization problems with a single objective, and you can also find in this book an extension to GA which is called non-dominated sorting genetic algorithm (NSGA) for solving multi-objective optimization problems. The book is available at Amazon and Springer:
- [Tutorial] Introduction to Optimization with Genetic Algorithm
- [Tutorial] Genetic Algorithm (GA) Optimization — Step-by-Step Example
Regarding the implementation of GA in Python, I also prepared a tutorial titled “Genetic Algorithm Implementation in Python” which discusses how to implement GA in details. It is available at these links:
https://www.linkedin.com/pulse/genetic-algorithm-implementation-python-ahmed-gad
There’s also a GitHub project that holds the Python implementation discussed in this tutorial, available here:
Genetic Algorithm Steps
GA is a random-based optimization technique that has a number of generic steps that are generally followed to solve any optimization problem. These steps are then customized to the problem being solved. This tutorial discusses these steps briefly but concentrates on how to customize them according to this project. The summary of these steps is as follows:
Data Representation
The first task for an optimization problem using GA is to think about the best way to represent the data. GA accepts the chromosome (i.e. solution) as a 1D row vector. The input image will not be 1D. The image may be 2D if it’s a binary or a gray image.
There may be more than 2 dimensions if the input image is color. If it’s RGB, for example, then there are 3 dimensions, one for each channel. Any data of more than 1 dimension must be represented in a 1D vector. How can we convert the multi-dimensional (MD) data into a 1D vector?
Starting with the simplest case in which the input is a 2D image (i.e. 2D matrix), converting it into a 1D vector requires us to merge the 2 dimensions into a single dimension. The matrix has multiple rows, and it’s necessary to merge all of these rows into a single row. This can be achieved by stacking the different rows together. This is illustrated in the next figure. The figure shows an image/matrix of 3 rows and 3 columns. That’s a total of 3×3=9 elements.
When converting to a 1D vector, the vector length will definitely be 9. The first 3 elements of this vector will be taken from the first row in the image. The next 3 elements of the vector will be taken from the second row, and finally the last 3 elements in the vector will be fetched from the last row.

After converting the 2D image into a 1D vector, it’s important to know how to restore the image back from the vector. This will be helpful in this project. In order to do that, it’s essential to know what the size of the image was. Knowing it was 3×3, the first 3 elements of the vector will form the first row in the image, the next 3 elements in the vector will form the next row, and so on.
According to the previous discussion, it’s now clear how to create the chromosome from the 2D image. Assuming the input image is not 2D but an RGB 3D color image, how do we do this conversion? This is also simple because the 3D RGB image can be visualized as 3 2D images—each image represents a channel of the RGB color space. So there is no single 2D image to convert but 3. The previous work of converting the 2D image into a 1D vector will just be repeated 3 times.
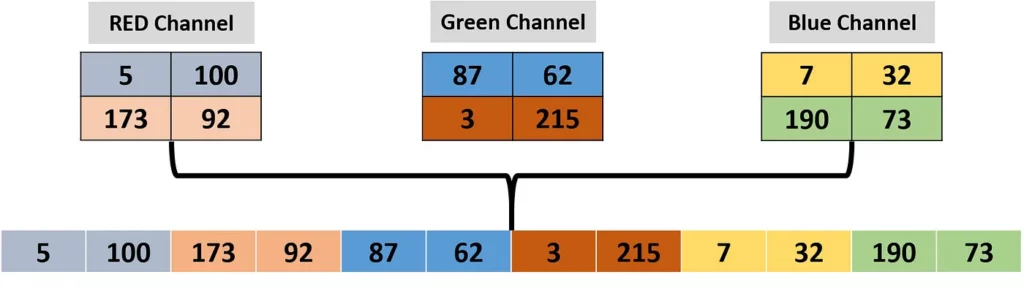
The next figure gives an example in which a 3D image (with 2 rows and 2 columns) is to be converted into a 1D row vector (i.e. chromosome). The 3D image is visualized individually, as 3 images in which each one is just a 2D image/matrix.
Because the resulting vector will hold all elements in all 3 2×2 images, then its length will be 3x2x2=12. The first image has 2×2=4 elements. These 4 elements are placed at the beginning of the vector. The 4 elements of the next image will be placed after these elements. Finally, the 4 elements of the last image will be placed at the end of the vector. Note that the project is not just limited to 3D images but can accept images in color spaces with more than 3 channels.
Converting the MD data into a 1D vector is the end for representing the data of this project, but it might not be the end for other types of problems. In this project, value encoding is used. The same values in the image are used in the chromosome. In other problems, it might be preferred to encode the values in different ways. In this case, there is an encoding step between the original form of the data and the chromosome. The encoding might be binary, octal, or hexadecimal, for example.
It’s also important to extract the MD image from the 1D vector. Given that the image has 3 channels, the vector will be divided into 3 parts of equal length. This length is equal to the number of elements within each channel. If the channel size is 2×2, then the first 4 elements of the vector will create the first channel in the image. The first 2 elements of these 4 will create the first row in this channel, and the last 2 elements of these 4 will be the second row for the same channel. The next 4 elements in the vector will create the next channel, and so on.
Python Code for Converting the Image into a Chromosome and Vice Versa
After understanding the concept well, we can build a Python function that accepts an image and returns its chromosome representation as a 1D row vector. The function is named img2chromosome() which is shown below. The function accepts the image to be converted as an argument named img_arr.
Using the reshape() function of the NumPy library, we can reshape any array from its current shape into a new shape. This function accepts the array and its new shape and returns an array of that shape:
In order to make the code independent on a specific image shape and able to work the same regardless of an image’s number of dimensions, the reduce() function from the functools library is used.
It accepts an operation and a list of numbers and applies this operation between every 2 numbers until returning just a single number. The operation it accepted is mul, which is short for multiplication. This operation is defined in the operator library. The list of numbers the reduce() function accepts is the shape of the image returned by the img_arr.shape property.
Assume that the shape is equal to (5, 6, 3). That is, the image has 3 channels and each channel has 5 rows and 6 columns. The reduce() function works by multiplying the first 2 numbers (5×6=30). The result is also multiplied by the third number to return 30×3=90. As a result, the input image will be reshaped into a vector of a single row, which consists of 90 columns.
The reverse of this process is also important. That is, converting that row vector into the original matrix. This can be implemented into another function named chromosome2img(), as listed below. This function accepts the chromosome and the image shape. By simply passing these arguments to the numpy.reshape() function, the original image can be restored:
At this time, the input image can be converted into a chromosome. and the image can be restored back from that chromosome.
After creating the chromosome, the next step is to build the initial population, which is simply a group of chromosomes.
Initial Population
GA starts an initial population, which is a group of solutions (chromosomes) to the given problem. These solutions are randomly generated. A Python function named initial_population() is created (as shown below) to return such a population.
Without looking at the function arguments, let’s think about what arguments are expected to be passed to it. At first, a chromosome (1D vector) is to be created. The vector length is equal to the number of elements in the image. Thus, there must be an argument to help in calculating this length. This is why the function accepts the shape of the image as the first argument named img_shape.
The function not only returns a single chromosome but a group of chromosomes. How many chromosomes should it return? It’s specified as the second argument n_individuals, which defaults to 8 if not specified:
Let’s assume that the image shape is (100, 50, 3). In other words, it’s an RGB image with 100 rows and 50 columns. Thus, the chromosome length is 15,000. If the number of chromosomes to return is 5, then there will be 5 chromosomes of length 15,000 each.
The first line in the function creates an empty NumPy array according to the number of chromosomes and their length. The remaining part fills these chromosomes by randomly generated numbers using the random() function inside the NumPy.random module.
Because these chromosomes/solutions are randomly generated, there is no guarantee that they will solve the problem correctly. This is why the GA evolves them using the following steps.
Fitness Calculation
GA starts with a number of bad solutions that are randomly generated. GA is based on the idea that evolving bad solutions might return better solutions. For every generation, the GA selects the best of the solutions in the current population and evolves them, hoping to return better solutions.
How does GA filter the solutions to return the best of them? This is done using a fitness function. Each chromosome is applied to this function and a number is returned. The higher the number, the better the solution. This is known as a maximization function.
The fitness function used in this project is named fitness_fun() and is listed below. It accepts 2 arguments—the target image and the current solution—and returns a number to measure the similarity between them:
At first, the mean of absolute differences between the elements in the target image and the current image are calculated. Based on this value, the lower the value, the better the solution. But the target is to return a value that is better when it’s higher. It’s better to do that in order to meet the specifications of GA. This is why it’s subtracted from the summation of all elements in the target image. In this way, the higher the value returned, the better the solution.
The previous function accepts a single solution and returns its fitness value. In order to calculate the fitness value for all solutions, a function named cal_pop_fitness() is implemented, as shown below. It simply loops through the solutions within the population.
For each solution, the previous fitness_fun() is called to return its fitness value. The fitness values for all solutions are saved in an array named qualities, which is finally returned by this function:
After calculating the fitness values for all solutions, the next step is to select the best of them for creating the next population. The best of these solutions are called parents. By mating these parents, the expected return is better solutions (offspring).
Parent Selection
The parents selected from a given population are the best solutions within it. When we say “best solutions”, we’re referring to the solutions with the highest fitness values.
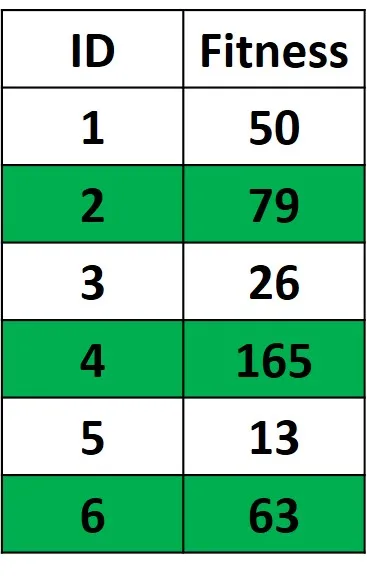
Assume that the population has 6 solutions and their fitness values are as given in the figure below. Before selecting the best parents, we need to decide how many parents to select. Assuming that half (3 out of 6) of the solutions will be selected, then the best 3 solutions are the ones with the highest fitness values. This why the solutions with ID 4, 2, and 6 are selected.
The parents are returned in this project according to a function named select_mating_pool() (shown below). It accepts 3 arguments: population (pop), the fitness values (qualities), and the number of parents (num_parents). It loops through the parents to select the ones with the highest fitness values and return them into an array named parents.
The function works by searching for the solution with the maximum fitness value and returning it into the parents array. In order to avoid selecting this parent again in the next iteration of the loop, its fitness value is set to -1. This guarantees not selecting it again. Then the next solution with the second maximum fitness value is selected, and the process repeats until returning all parents required:
After selecting the parents, our next step is to mate them for creating the new generation. Mating is applied using 2 operations: crossover and mutation.
Crossover
Mating 2 organisms means creating a new offspring that shares the genes inside both of them. The crossover operation selects a number of genes from each parent and places them into their offspring.
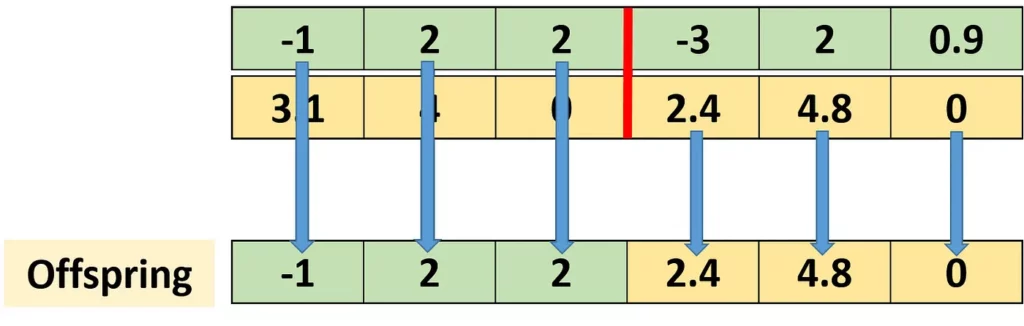
The next figure shows an example in which single-point crossover is applied between 2 parents in order to create an offspring. Single-point crossover works by selecting a point at the chromosome. Genes before this point are selected from one parent, and genes after it are selected from the other parent. As a result, the offspring shares genes from both parents.
The crossover operation is applied in the project using a function named crossover(). It accepts 3 arguments: the parents selected previously using the select_mating_pool() function, the input image shape (img_shape), and the number of offspring to return (n_individuals), which defaults to 8.
The function goes through the parents, selecting 2 of them for mating and producing an offspring. Then it moves to another 2 parents and repeats the process:
def crossover(parents, img_shape, n_individuals=8):
new_population = numpy.empty(shape=(n_individuals, functools.reduce(operator.mul, img_shape)), dtype=numpy.uint8)
#Previous parents (best elements).
new_population[0:parents.shape[0], :] = parents
# Getting how many offspring to be generated. If the population size is 8 and number of parents mating is 4, then number of offspring to be generated is 4.
num_newly_generated = n_individuals-parents.shape[0]
# Getting all possible permutations of the selected parents.
parents_permutations = list(itertools.permutations(iterable=numpy.arange(0, parents.shape[0]), r=2))
# Randomly selecting the parents permutations to generate the offspring.
selected_permutations = random.sample(range(len(parents_permutations)),
num_newly_generated)
comb_idx = parents.shape[0]
for comb in range(len(selected_permutations)):
# Generating the offspring using the permutations previously selected randmly.
selected_comb_idx = selected_permutations[comb]
selected_comb = parents_permutations[selected_comb_idx]
# Applying crossover by exchanging half of the genes between two parents.
half_size = numpy.int32(new_population.shape[1]/2)
new_population[comb_idx+comb, 0:half_size] = parents[selected_comb[0],
0:half_size]
new_population[comb_idx+comb, half_size:] = parents[selected_comb[1],
half_size:]
return new_populationLet’s assume that all solutions in a given population share a gene that represents a bad property. After applying crossover, this gene will definitely be available in the offspring. If the offspring takes its genes from 2 parents where each parent has a bad gene, then the offspring will now have 2 bad genes.
As a result, it will be worse than its parents. This is why it’s preferable that the next generation keep the previous parents in addition to the generated offspring. Even if the offspring are worse than the parents, the parents will be kept to avoid moving GA toward solutions that aren’t evolved. This is why the crossover() function returns the new population, which consists of both the current parents and their offspring.
But what do we do if the offspring is always worse than the parent? There is an operation called mutation that’s applied after crossover in order to introduce modifications to the offspring.
These modifications might solve a problem in the parents by replacing a bad gene with a better one. It might also introduce a problem by replacing a good gene and making it worse. If that happens, then the parents kept previously will prevent the selection of these bad offspring.
Mutation
The mutation operation selects some genes within the chromosome and then randomly changes their values.
It’s implemented according to the mutation() function listed below. It accepts the population returned by the crossover() function, number of parents, and the percent of the genes to be changed. The number of parents is passed in order to simply apply the mutation over the offspring and skip the parents.
Avoid setting the percentage to a high value because it will introduce many random changes, which will definitely lead to bad solutions. Slight changes using small percentages might (but not certainly) introduce good changes. This function returns the offspring after introducing such random changes:
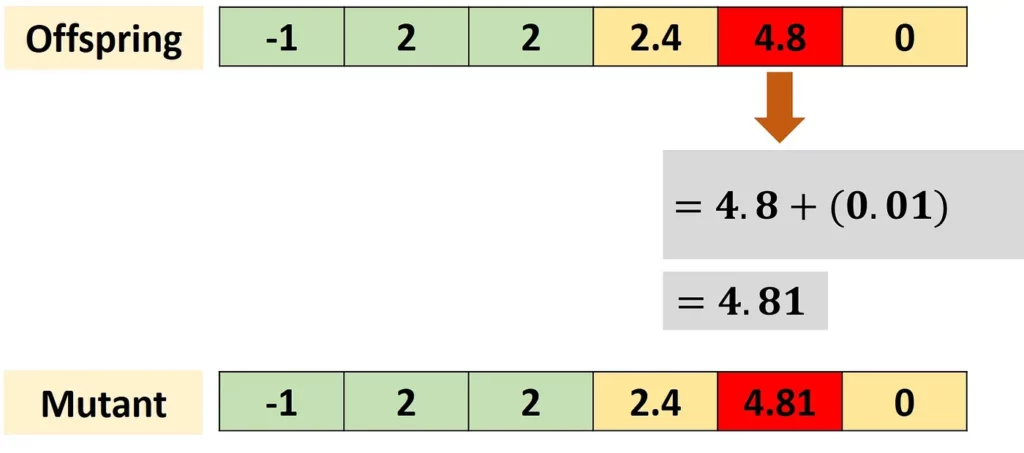
The previous figure applied the crossover and returned an offspring to which the mutation is applied, as shown in next figure. Assuming that a single gene is selected for mutation, its value is changed by randomly generating a number that is added to it.
The result of this addition will replace the previous gene value. The random change may be in other forms than addition. In this project, as implemented in the mutation() function, the random change is made by replacing the previous values with a new value within the range 0–255. This range is selected because the images are represented as 8-bit unsigned integers.
After the mutation operation is applied, the end of the current generation is reached and a new generation starts. The population used in this new generation is the combination between the parents selected from the previous generation in addition to the offspring returned after the mutation operation is applied.
Project Example
The project includes a file named Example_GeneticAlgorithm.py, which provides an example of how to run the project. At first, a color image is read and returned into a NumPy array. This image is available in the GitHub project. This image is then converted into a chromosome (1D vector) using the img2chromosome() function. In order to run the project, 3 parameters are specified:
- Population size: number of individuals per population
- Mating pool size: number of selected parents in the mating pool.
- Mutation percentage: number of genes whose values are changed in mutation.
After specifying these parameters, the initial population is created using the initial_population() function:
import os
import sys
import numpy
import scipy.misc
import itertools
import GARI
"""
Reproduce a single image using Genetic Algorithm (GA) by evolving
single pixel values.
This project works with both color and gray images without any modifications.
Just give the image path.
Using three parameters, we can customize it to statisfy our need.
The parameters are:
1) Population size. I.e. number of individuals pepr population.
2) Mating pool size. I.e. Number of selected parents in the mating pool.
3) Mutation percentage. I.e. number of genes to change their values.
Value encoding used for representing the input.
Crossover is applied by exchanging half of genes from two parents.
Mutation is applied by randomly changing the values of randomly selected
predefined percent of genes from the parents chromosome.
This project is implemented using Python 3.5 by Ahmed F. Gad.
Contact info:
[email protected]
https://www.linkedin.com/in/ahmedfgad/
"""
# Reading target image to be reproduced using Genetic Algorithm (GA).
target_im = scipy.misc.imread('fruit.jpg')
# Target image after enconding. Value encoding is used.
target_chromosome = GARI.img2chromosome(target_im)
# Population size
sol_per_pop = 8
# Mating pool size
num_parents_mating = 4
# Mutation percentage
mutation_percent = .01
"""
There might be inconsistency between the number of selected mating parents and
number of selected individuals within the population.
In some cases, the number of mating parents are not sufficient to
reproduce a new generation. If that occurred, the program will stop.
"""
num_possible_permutations = len(list(itertools.permutations(iterable=numpy.arange(0,
num_parents_mating), r=2)))
num_required_permutations = sol_per_pop-num_possible_permutations
if(num_required_permutations>num_possible_permutations):
print(
"n*Inconsistency in the selected populatiton size or number of parents.*"
"nImpossible to meet these criteria.n"
)
sys.exit(1)
# Creating an initial population randomly.
new_population = GARI.initial_population(img_shape=target_im.shape,
n_individuals=sol_per_pop)
for iteration in range(10000):
# Measing the fitness of each chromosome in the population.
qualities = GARI.cal_pop_fitness(target_chromosome, new_population)
print('Quality : ', numpy.max(qualities), ', Iteration : ', iteration)
# Selecting the best parents in the population for mating.
parents = GARI.select_mating_pool(new_population, qualities,
num_parents_mating)
# Generating next generation using crossover.
new_population = GARI.crossover(parents, target_im.shape,
n_individuals=sol_per_pop)
"""
Applying mutation for offspring.
Mutation is important to avoid local maxima. Avoiding mutation makes
the GA falls into local maxima.
Also mutation is important as it adds some little changes to the offspring.
If the previous parents have some common degaradation, mutation can fix it.
Increasing mutation percentage will degarde next generations.
"""
new_population = GARI.mutation(population=new_population,
num_parents_mating=num_parents_mating,
mut_percent=mutation_percent)
"""
Save best individual in the generation as an image for later visualization.
"""
GARI.save_images(iteration, qualities, new_population, target_im.shape,
save_point=500, save_dir=os.curdir+'//')
# Display the final generation
GARI.show_indivs(new_population, target_im.shape)Conclusion + 2 Helper Functions
This is the end of the essential work in this project. There are 2 other helper functions named save_images() and show_indivs(). The first one saves the solutions (images) on the disk in order to monitor the progress of the algorithm. The other displays all solutions after the algorithm completes its generations.
The progress of the algorithm when applied to an RGB image is given in the next figure.

For Contacting the Author
- E-mail: [email protected]
- LinkedIn: https://linkedin.com/in/ahmedfgad/
- KDnuggets: https://kdnuggets.com/author/ahmed-gad
- TowardsDataScience: https://towardsdatascience.com/@ahmedfgad
- GitHub: https://github.com/ahmedfgad

Comments 0 Responses